Measurement¶
Measurement modules produce measurements from either images or objects produced by prior modules.
MeasureColocalization¶
MeasureColocalization measures the colocalization and correlation between intensities in different images (e.g., different color channels) on a pixel-by-pixel basis, within identified objects or across an entire image.
Given two or more images, this module calculates the correlation & colocalization (Overlap, Manders, Costes’ Automated Threshold & Rank Weighted Colocalization) between the pixel intensities. The correlation / colocalization can be measured for entire images, or a correlation measurement can be made within each individual object. Correlations / Colocalizations will be calculated between all pairs of images that are selected in the module, as well as between selected objects. For example, if correlations are to be measured for a set of red, green, and blue images containing identified nuclei, measurements will be made between the following:
- The blue and green, red and green, and red and blue images.
- The nuclei in each of the above image pairs.
A good primer on colocalization theory can be found on the SVI website.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | YES | YES |
Measurements made by this module¶
- Correlation: The correlation between a pair of images I and J, calculated as Pearson’s correlation coefficient. The formula is covariance(I ,J)/[std(I ) × std(J)].
- Slope: The slope of the least-squares regression between a pair of images I and J. Calculated using the model A × I + B = J, where A is the slope.
- Overlap coefficient: The overlap coefficient is a modification of Pearson’s correlation where average intensity values of the pixels are not subtracted from the original intensity values. For a pair of images R and G, the overlap coefficient is measured as r = sum(Ri * Gi) / sqrt (sum(Ri*Ri)*sum(Gi*Gi)).
- Manders coefficient: The Manders coefficient for a pair of images R and G is measured as M1 = sum(Ri_coloc)/sum(Ri) and M2 = sum(Gi_coloc)/sum(Gi), where Ri_coloc = Ri when Gi > 0, 0 otherwise and Gi_coloc = Gi when Ri >0, 0 otherwise.
- Manders coefficient (Costes Automated Threshold): Costes’ automated threshold estimates maximum threshold of intensity for each image based on correlation. Manders coefficient is applied on thresholded images as Ri_coloc = Ri when Gi > Gthr and Gi_coloc = Gi when Ri > Rthr where Gthr and Rthr are thresholds calculated using Costes’ automated threshold method.
- Rank Weighted Colocalization coefficient: The RWC coefficient for a pair of images R and G is measured as RWC1 = sum(Ri_coloc*Wi)/sum(Ri) and RWC2 = sum(Gi_coloc*Wi)/sum(Gi), where Wi is Weight defined as Wi = (Rmax - Di)/Rmax where Rmax is the maximum of Ranks among R and G based on the max intensity, and Di = abs(Rank(Ri) - Rank(Gi)) (absolute difference in ranks between R and G) and Ri_coloc = Ri when Gi > 0, 0 otherwise and Gi_coloc = Gi when Ri >0, 0 otherwise. (Singan et al. 2011, BMC Bioinformatics 12:407).
MeasureGranularity¶
MeasureGranularity outputs spectra of size measurements of the textures in the image.
Image granularity is a texture measurement that tries to fit a series of structure elements of increasing size into the texture of the image and outputs a spectrum of measures based on how well they fit. Granularity is measured as described by Ilya Ravkin (references below). The size of the starting structure element as well as the range of the spectrum is given as input.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | YES | YES |
Measurements made by this module¶
- Granularity: The module returns one measurement for each instance of the granularity spectrum.
References¶
- Serra J. (1989) Image Analysis and Mathematical Morphology, Vol. 1. Academic Press, London
- Maragos P. “Pattern spectrum and multiscale shape representation”, IEEE Transactions on Pattern Analysis and Machine Intelligence, 11, N 7, pp. 701-716, 1989
- Vincent L. (2000) “Granulometries and Opening Trees”, Fundamenta Informaticae, 41, No. 1-2, pp. 57-90, IOS Press, 2000.
- Vincent L. (1992) “Morphological Area Opening and Closing for Grayscale Images”, Proc. NATO Shape in Picture Workshop, Driebergen, The Netherlands, pp. 197-208.
- Ravkin I, Temov V. (1988) “Bit representation techniques and image processing”, Applied Informatics, v.14, pp. 41-90, Finances and Statistics, Moskow, (in Russian)
MeasureImageAreaOccupied¶
MeasureImageAreaOccupied measures the total area in an image that is occupied by objects.
This module reports the sum of the areas and perimeters of the objects defined by one of the Identify modules, or the area of the foreground in a binary image. If the input image has a mask (for example, created by the MaskImage module), the measurements made by this module will take the mask into account by ignoring the pixels outside the mask.
You can use this module to measure the number of pixels above a given threshold if you precede it with thresholding performed by Threshold, and then select the binary image output by Threshold to be measured by this module.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | YES | YES |
See also¶
See also IdentifyPrimaryObjects, IdentifySecondaryObjects, IdentifyTertiaryObjects.
Measurements made by this module¶
- AreaOccupied/VolumeOccupied: The total area (2D) or volume (3D) occupied by the input objects or binary image.
- Perimeter/SurfaceArea The total length of the perimeter (2D) or surface area (3D) of the input objects/binary image.
- TotalArea/TotalVolume: The total pixel area (2D) or volume (3D) of the image that was subjected to measurement, excluding masked regions.
MeasureImageIntensity¶
MeasureImageIntensity measures several intensity features across an entire image (excluding masked pixels).
For example, this module will sum all pixel values to measure the total image intensity. You can choose to measure all pixels in the image or restrict the measurement to pixels within objects that were identified in a prior module. If the image has a mask, only unmasked pixels will be measured.
Note that for publication purposes, the units of intensity from microscopy images are usually described as “Intensity units” or “Arbitrary intensity units” because microscopes are not calibrated to an absolute scale. Also, it is important to note whether you are reporting the mean vs. the integrated intensity, so specify “Mean intensity units” or “Integrated intensity units” accordingly.
Keep in mind that the default behavior in CellProfiler is to rescale the image intensity from 0 to 1 by dividing all pixels in the image by the maximum possible intensity value. This “maximum possible” value is defined by the “Set intensity range from” setting in NamesAndTypes; see the help for that setting for more details.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | YES | YES |
See also¶
See also MeasureObjectIntensity, MaskImage.
Measurements made by this module¶
- TotalIntensity: Sum of all pixel intensity values.
- MeanIntensity, MedianIntensity: Mean and median of pixel intensity values.
- StdIntensity, MADIntensity: Standard deviation and median absolute deviation (MAD) of pixel intensity values. The MAD is defined as the median(|xi - median(x)|).
- MinIntensity, MaxIntensity: Minimum and maximum of pixel intensity values.
- LowerQuartileIntensity: The intensity value of the pixel for which 25% of the pixels in the object have lower values.
- UpperQuartileIntensity: The intensity value of the pixel for which 75% of the pixels in the object have lower values.
- TotalArea: Number of pixels measured, e.g., the area of the image excluding masked regions.
MeasureImageOverlap¶
MeasureImageOverlap calculates how much overlap occurs between the white portions of two black and white images
This module calculates overlap by determining a set of statistics that measure the closeness of an image to its true value. One image is considered the “ground truth” (possibly the result of hand-segmentation) and the other is the “test” image; the images are determined to overlap most completely when the test image matches the ground truth perfectly. The module requires binary (black and white) input, where the foreground of the images is white and the background is black. If you segment your images in CellProfiler using IdentifyPrimaryObjects, you can create such an image using ConvertObjectsToImage by selecting Binary as the color type. If your images have been segmented using other image processing software, or you have hand-segmented them in software such as Photoshop, you may need to use one or more of the following to prepare the images for this module:
- ImageMath: If the objects are black and the background is white, you must invert the intensity using this module.
- Threshold: If the image is grayscale, you must make it binary using this module, or alternately use an Identify module followed by ConvertObjectsToImage as described above.
- ColorToGray: If the image is in color, you must first convert it to grayscale using this module, and then use Threshold to generate a binary image.
In the test image, any foreground (white) pixels that overlap with the foreground of the ground truth will be considered “true positives”, since they are correctly labeled as foreground. Background (black) pixels that overlap with the background of the ground truth image are considered “true negatives”, since they are correctly labeled as background. A foreground pixel in the test image that overlaps with the background in the ground truth image will be considered a “false positive” (since it should have been labeled as part of the background), while a background pixel in the test image that overlaps with foreground in the ground truth will be considered a “false negative” (since it was labeled as part of the background, but should not be).
For 3D images, all image planes are concatenated into one large XY image and the overlap is computed on the transformed image.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | YES | YES |
Measurements made by this module¶
- True positive rate: Total number of true positive pixels / total number of actual positive pixels.
- False positive rate: Total number of false positive pixels / total number of actual negative pixels.
- True negative rate: Total number of true negative pixels / total number of actual negative pixels.
- False negative rate: Total number of false negative pixels / total number of actual positive pixels.
- Precision: Number of true positive pixels / (number of true positive pixels + number of false positive pixels).
- Recall: Number of true positive pixels/ (number of true positive pixels + number of false negative pixels).
- F-factor: 2 × (precision × recall)/(precision + recall). Also known as F1 score, F-score or F-measure.
- Earth mover’s distance: The minimum distance required to move each foreground pixel in the test image to some corresponding foreground pixel in the reference image.
- Rand index: A measure of the similarity between two data clusterings. Perfectly random clustering returns the minimum score of 0, perfect clustering returns the maximum score of 1.
- Adjusted Rand index: A variation of the Rand index which considers a correction for chance.
References¶
- Collins LM, Dent CW (1988) “Omega: A general formulation of the Rand Index of cluster recovery suitable for non-disjoint solutions”, Multivariate Behavioral Research, 23, 231-242. (link)
- Pele O, Werman M (2009) “Fast and Robust Earth Mover’s Distances”, 2009 IEEE 12th International Conference on Computer Vision.
MeasureImageQuality¶
MeasureImageQuality measures features that indicate image quality.
This module collects measurements indicating possible image aberrations, e.g., blur (poor focus), intensity, saturation (i.e., the percentage of pixels in the image that are at/near the maximum possible value, and at/near the minimum possible value). Details and guidance for each of these measures is provided in the settings help.
Please note that for best results, this module should be applied to the original raw images, rather than images that have already been corrected for illumination.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | YES | YES |
Measurements made by this module¶
Blur metrics
- FocusScore: A measure of the intensity variance across the image. This score is calculated using a normalized variance, which was the best-ranking algorithm for brightfield, phase contrast, and DIC images (Sun, 2004). Higher focus scores correspond to lower bluriness. More specifically, the focus score computes the intensity variance of the entire image divided by mean image intensity. Since it is tailored for autofocusing applications (difference focus for the same field of view), it assumes that the overall intensity and the number of objects in the image is constant, making it less useful for comparison images of different fields of view. For distinguishing extremely blurry images, however, it performs well.
- LocalFocusScore: A measure of the intensity variance between image sub-regions. A local version of the Focus Score, it subdivides the image into non-overlapping tiles, computes the normalized variance for each, and takes the mean of these values as the final metric. It is potentially more useful for comparing focus between images of different fields of view, but is subject to the same caveats as the Focus Score. It can be useful in differentiating good versus badly segmented images in the cases when badly segmented images usually contain no cell objects with high background noise.
- Correlation: A measure of the correlation of the image for a given spatial scale. This is a measure of the image spatial intensity distribution computed across sub-regions of an image for a given spatial scale (Haralick, 1973). If an image is blurred, the correlation between neighboring pixels becomes high, producing a high correlation value. A similar approach was found to give optimal performance for fluorescence microscopy applications (Vollath, 1987). Some care is required in selecting an appropriate spatial scale because differences in the spatial scale capture various features: moderate scales capture the blurring of intracellular features better than small scales and larger scales are more likely to reflect cell confluence than focal blur. You should select a spatial scale no bigger than the objects of interest, although you can select as many scales as desired and check empirically which is best.
- PowerLogLogSlope: The slope of the image log-log power spectrum. The power spectrum contains the frequency information of the image, and the slope gives a measure of image blur. A higher slope indicates more lower frequency components, and hence more blur (Field, 1997). This metric is recommended for blur detection in most cases.
Saturation metrics
- PercentMaximal: Percent of pixels at the maximum intensity value of the image.
- PercentMinimal: Percent of pixels at the minimum intensity value of the image.
Intensity metrics
- TotalIntensity: Sum of all pixel intensity values.
- MeanIntensity, MedianIntensity: Mean and median of pixel intensity values.
- StdIntensity, MADIntensity: Standard deviation and median absolute deviation (MAD) of pixel intensity values.
- MinIntensity, MaxIntensity: Minimum and maximum of pixel intensity values.
- TotalArea/TotalVolume: Number of pixels (or voxels) measured.
- Scaling: if Yes is chosen for “Include the image rescaling value?”, image’s rescaling value will be stored as a quality control metric. This is useful in confirming that all images are rescaled by the same value, given that some acquisition device vendors may output this value differently.
Threshold metrics:
Threshold: The automatically calculated threshold for each image for the thresholding method of choice.
The thresholds are recorded individually for each image and also as aggregate statistics for all images in the experiment. The mean, median and standard deviation of the threshold values across all images in the experiment are computed for each of the threshold methods selected and recorded as a measurement in the per-experiment table.
References¶
- Bray MA, Fraser AN, Hasaka TP, Carpenter AE (2012) “Workflow and metrics for image quality control in large-scale high-content screens.” J Biomol Screen 17(2):266-74. (link)
- Field DJ (1997) “Relations between the statistics of natural images and the response properties of cortical cells” Journal of the Optical Society of America. A, Optics, image science, and vision, 4(12):2379-94. (pdf)
- Haralick RM (1979) “Statistical and structural approaches to texture” Proc. IEEE, 67(5):786-804. (link)
- Vollath D (1987) “Automatic focusing by correlative methods” Journal of Microscopy 147(3):279-288. (link)
- Sun Y, Duthaler S, Nelson B (2004) “Autofocusing in computer microscopy: Selecting the optimal focus algorithm” Microscopy Research and Technique, 65:139-149 (link)
MeasureImageSkeleton¶
MeasureImageSkeleton measures the number of branches and endpoints in a skeletonized structure such as neurons, roots, or vasculature.
This module can analyze the number of total branches and endpoints for branching objects in an image. A branch is a pixel with more than two neighbors and an endpoint is a pixel with only one neighbor.
You can create a morphological skeleton with the MorphologicalSkeleton module from the Advanced category.
See also MeasureObjectSkeleton.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | YES | YES |
Measurements made by this module¶
- Branches: Total number of pixels with more than two neighbors.
- Endpoints: Total number of pixels with only one neighbor.
MeasureObjectIntensity¶
MeasureObjectIntensity measures several intensity features for identified objects.
Given an image with objects identified (e.g., nuclei or cells), this module extracts intensity features for each object based on one or more corresponding grayscale images. Measurements are recorded for each object.
Intensity measurements are made for all combinations of the images and objects entered. If you want only specific image/object measurements, you can use multiple MeasureObjectIntensity modules for each group of measurements desired.
Note that for publication purposes, the units of intensity from microscopy images are usually described as “Intensity units” or “Arbitrary intensity units” because microscopes are not calibrated to an absolute scale. Also, it is important to note whether you are reporting the mean vs. the integrated intensity, so specify “Mean intensity units” or “Integrated intensity units” accordingly.
Keep in mind that the default behavior in CellProfiler is to rescale the image intensity from 0 to 1 by dividing all pixels in the image by the maximum possible intensity value. This “maximum possible” value is defined by the “Set intensity range from” setting in NamesAndTypes; see the help for that setting for more details.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | YES | YES |
See also¶
See also NamesAndTypes, MeasureImageIntensity.
Measurements made by this module¶
- IntegratedIntensity: The sum of the pixel intensities within an object.
- MeanIntensity: The average pixel intensity within an object.
- StdIntensity: The standard deviation of the pixel intensities within an object.
- MaxIntensity: The maximal pixel intensity within an object.
- MinIntensity: The minimal pixel intensity within an object.
- IntegratedIntensityEdge: The sum of the edge pixel intensities of an object.
- MeanIntensityEdge: The average edge pixel intensity of an object.
- StdIntensityEdge: The standard deviation of the edge pixel intensities of an object.
- MaxIntensityEdge: The maximal edge pixel intensity of an object.
- MinIntensityEdge: The minimal edge pixel intensity of an object.
- MassDisplacement: The distance between the centers of gravity in the gray-level representation of the object and the binary representation of the object.
- LowerQuartileIntensity: The intensity value of the pixel for which 25% of the pixels in the object have lower values.
- MedianIntensity: The median intensity value within the object.
- MADIntensity: The median absolute deviation (MAD) value of the intensities within the object. The MAD is defined as the median(|xi - median(x)|).
- UpperQuartileIntensity: The intensity value of the pixel for which 75% of the pixels in the object have lower values.
- Location_CenterMassIntensity_X, Location_CenterMassIntensity_Y: The (X,Y) coordinates of the intensity weighted centroid (= center of mass = first moment) of all pixels within the object.
- Location_MaxIntensity_X, Location_MaxIntensity_Y: The (X,Y) coordinates of the pixel with the maximum intensity within the object.
MeasureObjectIntensityDistribution¶
MeasureObjectIntensityDistribution measures the spatial distribution of intensities within each object.
Given an image with objects identified, this module measures the intensity distribution from each object’s center to its boundary within a set of bins, i.e., rings that you specify.
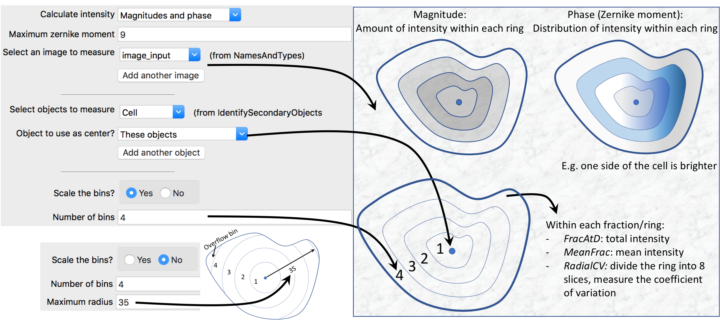
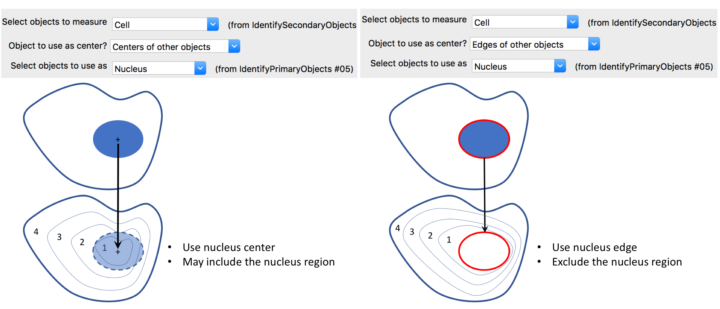
The distribution is measured from the center of the object, where the center is defined as the point farthest from any edge. The numbering of bins is from 1 (innermost) to N (outermost), where N is the number of bins you specify. Alternatively, if primary objects exist within the object of interest (e.g., nuclei within cells), you can choose the center of the primary objects as the center from which to measure the radial distribution. This might be useful in cytoplasm-to-nucleus translocation experiments, for example. Note that the ring widths are normalized per-object, i.e., not necessarily a constant width across objects.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | NO | YES |
See also¶
See also MeasureObjectIntensity and MeasureTexture.
Measurements made by this module¶
- FracAtD: Fraction of total stain in an object at a given radius.
- MeanFrac: Mean fractional intensity at a given radius; calculated as fraction of total intensity normalized by fraction of pixels at a given radius.
- RadialCV: Coefficient of variation of intensity within a ring, calculated across 8 slices.
- Zernike: The Zernike features characterize the distribution of intensity across the object. For instance, Zernike 1,1 has a high value if the intensity is low on one side of the object and high on the other. The ZernikeMagnitude feature records the rotationally invariant degree magnitude of the moment and the ZernikePhase feature gives the moment’s orientation.
MeasureObjectNeighbors¶
MeasureObjectNeighbors calculates how many neighbors each object has and records various properties about the neighbors’ relationships, including the percentage of an object’s edge pixels that touch a neighbor.
Given an image with objects identified (e.g., nuclei or cells), this module determines how many neighbors each object has. You can specify the distance within which objects should be considered neighbors, or that objects are only considered neighbors if they are directly touching.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | YES | NO |
See also¶
See also the Identify modules.
Measurements made by this module¶
Object measurements
- NumberOfNeighbors: Number of neighbor objects.
- PercentTouching: Percent of the object’s boundary pixels that touch neighbors, after the objects have been expanded to the specified distance. Note that this measurement is only available if you use the same set of objects for both objects and neighbors.
- FirstClosestObjectNumber: The index of the closest object.
- FirstClosestDistance: The distance to the closest object (in units of pixels).
- SecondClosestObjectNumber: The index of the second closest object.
- SecondClosestDistance: The distance to the second closest object (in units of pixels).
- AngleBetweenNeighbors: The angle formed with the object center as the vertex and the first and second closest object centers along the vectors.
Object relationships: The identity of the neighboring objects, for each object. Since per-object output is one-to-one and neighbors relationships are often many-to-one, they may be saved as a separate file in ExportToSpreadsheet by selecting Object relationships from the list of objects to export.
Technical notes¶
Objects discarded via modules such as IdentifyPrimaryObjects or IdentifySecondaryObjects will still register as a neighbors for the purposes of accurate measurement. For instance, if an object touches a single object and that object had been discarded, NumberOfNeighbors will be positive, but there will not be a corresponding ClosestObjectNumber.
MeasureObjectSizeShape¶
MeasureObjectSizeShape measures several area and shape features of identified objects.
Given an image with identified objects (e.g., nuclei or cells), this module extracts area and shape features of each one. Note that these features are only reliable for objects that are completely inside the image borders, so you may wish to exclude objects touching the edge of the image using Identify settings for 2D objects, or by applying FilterObjects downstream.
The display window for this module shows per-image aggregates for the per-object measurements. If you want to view the per-object measurements themselves, you will need to use an Export module to export them, or use DisplayDataOnImage to display the object measurements of choice overlaid on an image of choice.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | YES | NO |
See also¶
See also MeasureImageAreaOccupied.
Measurements made by this module¶
Some measurements are available for 3D and 2D objects, while some are 2D only.
See the Technical Notes below for an explanation of a key step underlying many of the following metrics: creating an ellipse with the same second-moments as each object.
Area: (2D only) The number of pixels in the region.
Volume: (3D only) The number of voxels in the region.
Perimeter: (2D only) The total number of pixels around the boundary of each region in the image.
SurfaceArea: *(3D only)# The total number of voxels around the boundary of each region in the image.
FormFactor: (2D only) Calculated as 4*π*Area/Perimeter2. Equals 1 for a perfectly circular object.
Solidity: (2D only) The proportion of the pixels in the convex hull that are also in the object, i.e., ObjectArea/ConvexHullArea.
Extent: The proportion of the pixels (2D) or voxels (3D) in the bounding box that are also in the region. Computed as the area/volume of the object divided by the area/volume of the bounding box.
EulerNumber: (2D only) The number of objects in the region minus the number of holes in those objects, assuming 8-connectivity.
Center_X, Center_Y, Center_Z: The x-, y-, and (for 3D objects) z- coordinates of the point farthest away from any object edge (the centroid). Note that this is not the same as the Location-X and -Y measurements produced by the Identify or Watershed modules or the Location-Z measurement produced by the Watershed module.
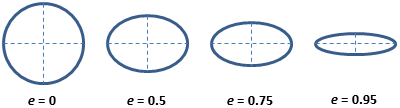
Eccentricity: (2D only) The eccentricity of the ellipse that has the same second-moments as the region. The eccentricity is the ratio of the distance between the foci of the ellipse and its major axis length. The value is between 0 and 1. (0 and 1 are degenerate cases; an ellipse whose eccentricity is 0 is actually a circle, while an ellipse whose eccentricity is 1 is a line segment.)
MajorAxisLength: (2D only) The length (in pixels) of the major axis of the ellipse that has the same normalized second central moments as the region.
MinorAxisLength: (2D only) The length (in pixels) of the minor axis of the ellipse that has the same normalized second central moments as the region.
Orientation: (2D only) The angle (in degrees ranging from -90 to 90 degrees) between the x-axis and the major axis of the ellipse that has the same second-moments as the region.
Compactness: (2D only) The mean squared distance of the object’s pixels from the centroid divided by the area. A filled circle will have a compactness of 1, with irregular objects or objects with holes having a value greater than 1.
MaximumRadius: (2D only) The maximum distance of any pixel in the object to the closest pixel outside of the object. For skinny objects, this is 1/2 of the maximum width of the object.
MedianRadius: (2D only) The median distance of any pixel in the object to the closest pixel outside of the object.
MeanRadius: (2D only) The mean distance of any pixel in the object to the closest pixel outside of the object.
MinFeretDiameter, MaxFeretDiameter: (2D only) The Feret diameter is the distance between two parallel lines tangent on either side of the object (imagine taking a caliper and measuring the object at various angles). The minimum and maximum Feret diameters are the smallest and largest possible diameters, rotating the calipers along all possible angles.
Zernike shape features: (2D only) These metrics of shape describe a binary object (or more precisely, a patch with background and an object in the center) in a basis of Zernike polynomials, using the coefficients as features (Boland et al., 1998). Currently, Zernike polynomials from order 0 to order 9 are calculated, giving in total 30 measurements. While there is no limit to the order which can be calculated (and indeed you could add more by adjusting the code), the higher order polynomials carry less information.
Technical notes¶
A number of the object measurements are generated by creating an ellipse with the same second-moments as the original object region. This is essentially the best-fitting ellipse for a given object with the same statistical properties. Furthermore, they are not affected by the translation or uniform scaling of a region.
Following computer vision conventions, the origin of the X and Y axes is at the top left of the image rather than the bottom left; the orientation of objects whose topmost point is on their right (or are rotated counter-clockwise from the horizontal) will therefore have a negative orientation, while objects whose topmost point is on their left (or are rotated clockwise from the horizontal) will have a positive orientation.
The Zernike features are computed within the minimum enclosing circle of the object, i.e., the circle of the smallest diameter that contains all of the object’s pixels.
References¶
- Rocha L, Velho L, Carvalho PCP, “Image moments-based structuring and tracking of objects”, Proceedings from XV Brazilian Symposium on Computer Graphics and Image Processing, 2002. (pdf)
- Principles of Digital Image Processing: Core Algorithms (Undergraduate Topics in Computer Science): Section 2.4.3 - Statistical shape properties
- Chrystal P (1885), “On the problem to construct the minimum circle enclosing n given points in a plane”, Proceedings of the Edinburgh Mathematical Society, vol 3, p. 30
MeasureObjectSkeleton¶
MeasureObjectSkeleton measures information for any branching structures, such as neurons, root or branch systems, vasculature, or any skeletonized system that originates from a single point (such as neurites branching from a single nucleus/soma).
This module measures the number of trunks and branches for each branching system in an image. The module takes a skeletonized image of the object plus previously identified seed objects (for instance, each neuron’s soma) and finds the number of axon or dendrite trunks that emerge from the soma and the number of branches along the axons and dendrites. Note that the seed objects must be both smaller than the skeleton, and touching the skeleton, in order to be counted.
The typical approach for this module is the following:
- Identify a seed object. This object is typically a nucleus, identified with a module such as IdentifyPrimaryObjects.
- Identify a larger object that touches or encloses this seed object. For example, the neuron cell can be grown outwards from the initial seed nuclei using IdentifySecondaryObjects.
- Use the Morph module to skeletonize the secondary objects.
- Finally, the primary objects and the skeleton objects are used as inputs to MeasureObjectSkeleton.
The module determines distances from the seed objects along the axons and dendrites and assigns branchpoints based on distance to the closest seed object when two seed objects appear to be attached to the same dendrite or axon.
The module records vertices which include trunks, branchpoints, and endpoints.
Note that this module was referred to as MeasureNeurons in previous versions of CellProfiler.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | NO | NO |
See also¶
See also MeasureImageSkeleton.
Measurements made by this module¶
- NumberTrunks: The number of trunks. Trunks are branchpoints that lie within the seed objects
- NumberNonTrunkBranches: The number of non-trunk branches. Branches are the branchpoints that lie outside the seed objects.
- NumberBranchEnds: The number of branch end-points, i.e, termini.
- TotalObjectSkeletonLength: The length of all skeleton segments per object.
MeasureTexture¶
MeasureTexture measures the degree and nature of textures within images and objects to quantify their roughness and smoothness.
This module measures intensity variations in grayscale images. An object or entire image without much texture has a smooth appearance; an object or image with a lot of texture will appear rough and show a wide variety of pixel intensities.
Note that any input objects specified will have their texture measured against all input images specified, which may lead to image-object texture combinations that are unnecessary. If you do not want this behavior, use multiple MeasureTexture modules to specify the particular image-object measures that you want.
Note also that CellProfiler in all 2.X versions increased speed by binning the image into only 8 greyscale levels before calculating Haralick features; this is not done in CellProfiler versions 3.0.0 and after. Values calculated in MeasureTexture in CellProfiler 2 versions will therefore not directly correspond to those in CellProfiler 3 and after.
| Supports 2D? | Supports 3D? | Respects masks? |
|---|---|---|
| YES | YES | YES |
Measurements made by this module¶
Haralick Features: Haralick texture features are derived from the co-occurrence matrix, which contains information about how image intensities in pixels with a certain position in relation to each other occur together. MeasureTexture can measure textures at different scales; the scale you choose determines how the co-occurrence matrix is constructed. For example, if you choose a scale of 2, each pixel in the image (excluding some border pixels) will be compared against the one that is two pixels to the right.
Thirteen measurements are then calculated for the image by performing mathematical operations on the co-occurrence matrix (the formulas can be found here):
- AngularSecondMoment: Measure of image homogeneity. A higher value of this feature indicates that the intensity varies less in an image. Has a value of 1 for a uniform image.
- Contrast: Measure of local variation in an image, with 0 for a uniform image and a high value indicating a high degree of local variation.
- Correlation: Measure of linear dependency of intensity values in an image. For an image with large areas of similar intensities, correlation is much higher than for an image with noisier, uncorrelated intensities. Has a value of 1 or -1 for a perfectly positively or negatively correlated image, respectively.
- Variance: Measure of the variation of image intensity values. For an image with uniform intensity, the texture variance would be zero.
- InverseDifferenceMoment: Another feature to represent image contrast. Has a low value for inhomogeneous images, and a relatively higher value for homogeneous images.
- SumAverage: The average of the normalized grayscale image in the spatial domain.
- SumVariance: The variance of the normalized grayscale image in the spatial domain.
- SumEntropy: A measure of randomness within an image.
- Entropy: An indication of the complexity within an image. A complex image produces a high entropy value.
- DifferenceVariance: The image variation in a normalized co-occurrence matrix.
- DifferenceEntropy: Another indication of the amount of randomness in an image.
- InfoMeas1: A measure of the total amount of information contained within a region of pixels derived from the recurring spatial relationship between specific intensity values.
- InfoMeas2: An additional measure of the total amount of information contained within a region of pixels derived from the recurring spatial relationship between specific intensity values. It is a complementary value to InfoMeas1 and is on a different scale.
Technical notes¶
To calculate the Haralick features, MeasureTexture normalizes the co-occurrence matrix at the per-object level by basing the intensity levels of the matrix on the maximum and minimum intensity observed within each object. This is beneficial for images in which the maximum intensities of the objects vary substantially because each object will have the full complement of levels.